Complexity and diversity of cellular systems
Biochemistry is necessary in order to understand living organisms: namely to appreciate their diversity, to study their similarities, and to understand their evolution. The following will give an overview over the complexity and diversity of cellular systems and will outline the importance to understand biochemical mechanisms in order to understand the mechanism of life at a cellular level.
Physics and chemistry are necessary for the understanding
of biochemistry which is a multi-disciplinary science. It interrelates
with biophysics, molecular biology, cell biology
and structural biology. The central domain of biochemistry is enzymology
which includes metabolic pathways, catalysis, and the structure-function
relationship of enzymes - the proteins that catalyze metabolic reactions.
| Field of research | Activities |
| ___________________ | ______________________________________________ |
| Biochemists | identify, isolate, and purify enzymes and characterize their functions |
| Molecular biologists | characterize the gene that codes for these proteins and map the chromosomal location of genes (chromosome organization) |
| Cell biologists | study the cellular localization of proteins and their interaction with other cell components |
| Biophysicists | and structural biologist measure the physical characteristics and structure of proteins |
A scientist who studies a protein must have a framework that includes a working hypothesis, a technique, and a theory. If one isolates a protein, one has to keep in mind where and when it has been isolated and what is already known about the cellular environment, the chemistry of the molecule's components and the function or genetic structure of the organism from which it has been isolated. In short, a protein, whatever function or structure one studies, can only be understood within the proper framework of the hierarchy of the system it belongs to. Understanding the hierarchical structure of the function and structure of biological system is key to the understanding of their physiology, development, and evolution (Voet&Voet, Fig. 1-14).
Fig. Hierarchical organization of bio-sciences
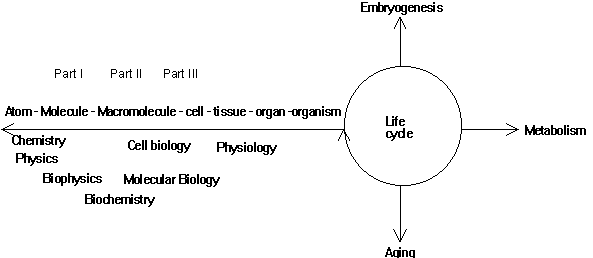
Biochemistry deals with but a little aspect of life. However, it deals
with the level of organic macromolecules, the building blocks of the cells,
which are the building blocks of tissues, organs, and organisms, which
are the building blocks of populations, societies and finally the ecosystems
of our world. The macromolecules themselves are build up from polymers,
which arise from monomers, which are made from atoms.
The synthesis and degradation of the monomer is the topic of metabolic biochemistry, and will not be covered in this course. However, the structure of several enzymes which are involved in metabolic pathways will be studied. It is necessary, therefore, to know the basics of the monomer, their chemistry and physics in solution.
Table Major biological macromolecules and their components
| Polymer | Monomer |
| protein | 20 amino acids |
| DNA | 4 nucleotides |
| polysaccharide | > 8 sugars |
| membrane | > 4 Lipids |
The table summarizes the contents of Part I of this course: how macromolecules or polymers are formed and which building blocks (monomers) they are made from. How, through the virtue of free combination, an astounding degree of complexity can be achieved both in terms of structure and function of the polymers. While a polymer is composed of many subunits that are covalently linked with each other, there exist molecular entities that exhibit larger complexes through non-covalent interactions. These supra-molecular structures are self-assembly systems constituting the basis of all living matter.
The biosynthesis of the polymer is of central importance in life. It is responsible for the transformation and accuracy of molecular structures and function that are stored in the genes and passed down from generation to generation of any given organism. The inheritance of structure and the accuracy of the process of polymer synthesis in the cell is absolutely important for the propagation of life and this process is the basis of Darwinian evolution. Being central to all life forms, the molecular components of transcription (DNA to RNA synthesis) and translation (protein biosynthesis using a mRNA template) are amongst the most conserved proteins among all organisms.
This is a central theme of Structural Biochemistry - to understand through factual and methodical knowledge how macromolecular structures relate to their function, and how the function of individual macromolecules relates to the working of an organism through a multitude of interactions.
We can ask many questions of what an organism is, what distinguishes
it from other organisms, what makes it alive. Let's first consider the
composition of a uni-cellular organism, the bacterium Escherichia coli.
| Component | % (weight) |
| H20 | 70 |
| Protein | 15 |
| Nucleic acid : DNA | 1 |
| Nucleic acid : RNA | 6 |
| Polysaccharides and precursors | 3 |
| Lipids and precursors | 2 |
| Other small organic molecules | 1 |
| Inorganic ions | 1 |
This list is a compilation of how much there is from which kind of chemicals.
Similar lists can be generated for plants and humans, fungi and insects.
The lists are quite comparable and any differences in ratios, if they
exist, cannot tell us much about the differences of the organisms compared,
but there are clear differences to the composition of inorganic matter.
| Earth's crust | % | Human | % | Pumpkin | % |
| Oxygen | 46.6 | Oxygen | 65 | Oxygen | 85 |
| Silicon | 27.7 | Carbon | 18 | Hydrogen | 10.7 |
| Aluminum | 8.1 | Hydrogen | 10 | Carbon | 3.3 |
| Iron | 5.0 | Nitrogen | 3 | Potassium | 0.34 |
| Calcium | 3.6 | Calcium | 2 | Nitrogen | 0.16 |
| Sodium | 2.8 | Phosphorus | 1.1 | Phosphorus | 0.05 |
| Potassium | 2.6 | Potassium | 0.35 | Calcium | 0.02 |
| Magnesium | 2.1 | Sulfur | 0.25 | Magnesium | 0.01 |
| Others | 1.5 | Sodium | 0.15 | Iron | 0.008 |
| - | - | Chlorine | 0.15 | Sodium | 0.001 |
| - | - | Magnesium | 0.05 | Zinc | 0.0002 |
| - | - | Iron | 0.004 | Copper | 0.0001 |
| - | - | Iodine | 0.0004 | Other | 0.00005 |
Obviously, and it might sound trivial, the organization of living matter in space and time (the development of an organism) are the important biological data to study. Considering all the variability in form and function encountered in nature, one might as a designer ask, how little it would take to build a viable organism. In other words:
- 1,830,137 base pairs- 1743 genes, or open reading frames, of which
- 736 open reading frames have not been assigned to any known function
- G+C content 38%, similar to that of human
During the past years the complete genome sequences of twenty-one microorganisms
have been completed and 63 more in progress (April 1998, see MAGPIE home
page http://www-fp.mcs.anl.gov/~gaasterland/magpie.html to find a running
list of genome projects; an alternative site for information on eukaryotic
organsims only is the genome monitoring table at http://www.ebi.ac.uk/~sterk/genome-MOT/index.html).
The first six organisms include the eubacteria Mycoplasma genitalium,
the archae Methanococcus jannaschii, the yeast Saccharomyces
cerevisiae, the Gram-negative eubacteria Escherichia coli K-12
and Helicobacter pylori representing the genomes from all three
domains of life: eubacteria, archaea, and eukarya. The first animal genome,
of the worm C. elegans, is expected to be completed this year.
The bacteria M.genitalium is a parasite and facultative pathogen of human genital and respiratory tracts and is even smaller than H. influenzae, because it relies heavily on the host cell metabolism. The circular genome contains 580,010 base pairs. With 12% non-coding regions it includes 470 predicted protein coding regions [Fraser et. al., 1995].
The first complete genome of a eukaryotic cell, the yeast S.cerevisiae, contains 16 chromosomes with a total of 12.06 million base pairs [Williams, N., 1996].
The archaeon M.jannaschii's complete genome contains a circular chromosome with 1.66 million base pairs plus two extra-chromosomal elements of 58kb and 16kb, with a total of 1738 predicted protein-coding genes, of which only 38% could be identified as putative enzymes with known cellular function (see section 3.9). M.jannaschii lives on the sea floor some 2.6km below the surface in the east Pacific and is able to grow at a pressure of 200atm, and a temperature range of 48-94° C, with the optimum at 85° C. As a strict anaerobe, it is an auxotrophic organism which can synthesize all cellular components from inorganic precursors (producing methane as byproduct, hence the name) [Buld, C.J., 1996].
The eubacteria Escherichia coli is a key organism for biological research. The completion of the sequencing of its 4.64Mb genome has been accomplished by two groups independently (see C. OBrien, Nature (1997) 385:472 for a comment). The complete genome can be found at the University of Wisconsin (http://www.genetics.wisc.edu/) and all organisms listed here can be found in the genome database of the National Center for Biotechnology Information (NCBI).
The gastric pathogen Helicobacter pylori has a circular genome of 1,667,867 base pairs and 1,590 predicted coding sequences [Tomb et al., 1997]. H.pylori is probably the most common chronical bacterial infection in humans: atrophic gastritis and peptic ulceration. It colonizes an environment of low pH (~2) and survives probably due to its ability to establish a positive inside membrane potential (all known cells have negative inside) and containing a high percentage of the basic amino acids arginine and lysine.
In addition, and years earlier (to give proper credit), several viral genomes have been sequenced: bacteriophage f X174 (5,386bp), in 1977, bacteriophage l (48,502bp), cytomegalovirus CMV (229kb), and Vaccinia (192kb). Note that, although viruses constitute replicating entities and can survive sometimes for long periods of time outside a host cell, their life cycle is strictly dependent on a host organism which provides the virus with the cellular components necessary for the replication of its genome and synthesis of coat structure: proteins and cell membrane (see section 3.8).
"If genes just make proteins and our proteins are the same, then why are we so different?"The minimal genetic requirement to sustain a living organism depends on the self-organizational properties of the macromolecules synthesized by the cell. The self-assembly process is critically dependent on the physical state of the system, i.e., the thermodynamic properties of the system and thus the equilibrium and non-equilibrium conditions of solutes in a solvent.
[Penman, S., 1991]
'Much of biological research practiced today is grounded in the chemistry that occurs in solution' (Penman). And much of biochemistry will be dealing with this chemistry in solution. The solvent is of paramount importance and the structure and function of proteins and nucleic acids in solution are well understood. This is the domain of biochemistry. Traditionally biochemistry dealt with the chemical equilibrium, but little or nothing in a living cell happens at the chemical equilibrium. Solutions, in addition, provide homogenous mixtures of cell components, but cell components are highly organized within the cell.
A possible answer to Penman's question is that 'proteins are the same' when studied in diluted aqueous solution. We are different in our physical organization. Time and space in the organization of macro-molecules of an organism determines its morphology. The embryonic development is an extremely precise mechanism of putting cellular components together at the right place and at the right time. 'Right' that is to say with respect to what we expect to see: offspring that resembles the parents. Evolution is the change in physical organization that is encoded by our genes and that can be expressed again and again. It is not possible to say that form of an organism is stored in its genes. Form is the result of sequential activities of different gene products (proteins). This sequential putting-together of molecules to form a cell leads to variability in our morphological traits because the putting-together itself is variable. The area of morphogenesis is a true challenge for today's biochemists and I hope to be able to show the fascination of what biochemistry can teach us in order to understand the nature of life.